Changing activations in spiking layers
Contents
Changing activations in spiking layers#
Sometimes it’s helpful to change spiking, membrane reset or backpropagation behaviour. Let’s look at a few examples based on the Integrate and Fire (IAF) layer.
import torch
import torch.nn as nn
import sinabs
import sinabs.layers as sl
import sinabs.activation as sina
import matplotlib.pyplot as plt
import numpy as np
We define a utility plotting function.
def plot_evolution(neuron_model: sinabs.layers, input: torch.Tensor):
neuron_model.reset_states()
v_mem = []
spikes = []
for step in range(input.shape[1]):
output = neuron_model(input[:, step])
v_mem.append(neuron_model.v_mem)
spikes.append(output)
plt.figure(figsize=(10, 3))
v_mem = torch.cat(v_mem).detach().numpy()
plt.plot(v_mem, drawstyle="steps", label="v_mem")
spikes = torch.cat(spikes).detach().numpy()
plt.plot(spikes, label="output", drawstyle="steps", color="black")
plt.xlabel("time")
plt.title(
f"{neuron_model.__class__.__name__} neuron dynamics with {neuron_model.spike_fn.__name__} and {neuron_model.reset_fn.__class__.__name__}."
)
plt.legend()
Single or Multi spike#
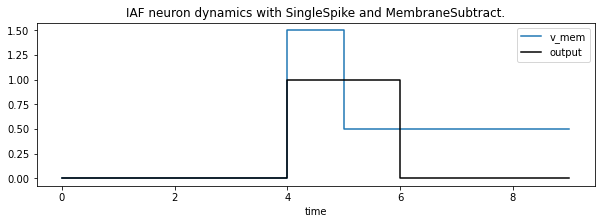
Let’s start by injecting a current 2.5 higher than the spike threshold. If the neuron uses a SingleSpike spike function, only a single spike can be emitted per time step. In combination with a MembraneSubtract reset function, which subtracts the spiking threshold for each spike, the neuron emits 2 spikes over 2 time steps and then is left with the remaining 0.5 as membrane potential.
iaf_neuron = sl.IAF(
spike_threshold=1.0, spike_fn=sina.SingleSpike, reset_fn=sina.MembraneSubtract()
)
single_current = torch.zeros((1, 10, 1))
single_current[:, 5] = 2.5
plot_evolution(iaf_neuron, single_current)
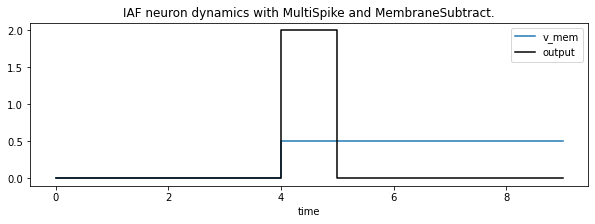
Here is the same input but now we changed the spike function to MultiSpike. The two spikes will now be emitted within a single time step. What would be the motivation for multiple spikes per time step? When discretizing a continuous input in time, the assumption of a single spike per time bin might get inaccurate. When reducing the number of time steps, potentially multiple spikes per bin reduces temporal quantisation error.
iaf_neuron = sl.IAF(
spike_threshold=1.0, spike_fn=sina.MultiSpike, reset_fn=sina.MembraneSubtract()
)
plot_evolution(iaf_neuron, single_current)
Membrane reset instead of subtract#
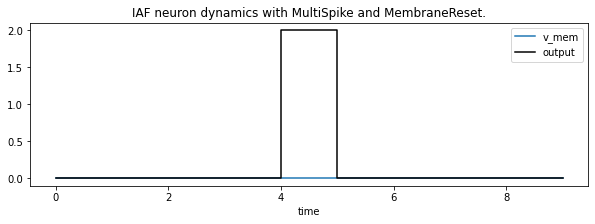
Alternatively we can also reset the membrane potential to 0 after each spike, no matter the output. Keep in mind that the input current is still 2.5 times the spike threshold so we’ll lose some information by doing that.
iaf_neuron = sl.IAF(
spike_threshold=1.0, spike_fn=sina.MultiSpike, reset_fn=sina.MembraneReset()
)
plot_evolution(iaf_neuron, single_current)
Surrogate gradient functions#
What follows is a very brief demonstration of surrogate gradients in SNNs. We feed a constant input current to a single neuron with a trainable weight and get a number of output spikes.
const_current = torch.ones((1, 100, 1)) * 0.03
torch.manual_seed(12345)
neuron = nn.Sequential(
nn.Linear(1, 1, bias=False),
sl.IAF(spike_threshold=1.0),
)
print(f"Sum of spikes: {neuron(const_current).sum()}")
Sum of spikes: 2.0
We define a loss function and a target sum of spikes of 5. This convex function is well suited for modern day optimizers.
criterion = nn.functional.mse_loss
target_firing_rate = torch.tensor(5.0)
losses = [criterion(target_firing_rate, torch.tensor(i)) for i in range(11)]
plt.plot(losses)
plt.title("Loss function for target firing rate 5")
plt.ylabel("Loss")
plt.xlabel("Output firing rate");
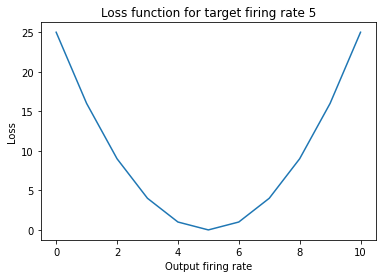
Now we sweep the loss with respect to different weights. You see that there are ranges of weight values that all have the same loss value, because the output of the network with its spike activation is highly non-linear. The gradients of this function unfortunately are mostly zero, which makes gradient descent difficult in this scenario.
losses = []
weights = torch.linspace(0, 3, 1000)
for w in weights:
neuron[0].weight = nn.Parameter(w.unsqueeze(0).unsqueeze(0))
neuron[1].reset_states()
output_sum = neuron(const_current).sum()
losses.append(criterion(target_firing_rate, output_sum).item())
plt.plot(weights, losses, label="loss")
plt.plot(weights, np.gradient(losses), label="derivative of loss")
plt.title("Loss function with respect to neuron weight.")
plt.xlabel("weight value of our IAF neuron")
plt.legend();
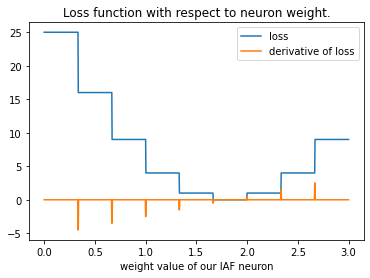
The gradients are zero nearly everywhere, which makes it very difficult to optimise. The reason for this is the non-linear activation function of each spiking neuron, which is a function of membrane potential. In the case of SingleSpike, it is the Heaviside function.
v_mem = torch.linspace(-0.5, 3.5, 500, requires_grad=True)
spike_threshold = 1.0
activations_singlespike = sina.SingleSpike.apply(
v_mem, spike_threshold, sina.MultiGaussian()
)
activations_multispike = sina.MultiSpike.apply(
v_mem, spike_threshold, sina.MultiGaussian()
)
plt.plot(
v_mem.detach(), activations_singlespike.detach(), label="SingleSpike activation fn"
)
plt.plot(
v_mem.detach(), activations_multispike.detach(), label="MultiSpike activation fn"
)
plt.plot(
v_mem.detach(),
np.gradient(activations_singlespike.detach()),
label="derivative of SingleSpike",
)
plt.title("Output as a function of membrane potential.")
plt.xlabel("Membrane potential v_mem")
plt.ylabel("output activation")
plt.legend();
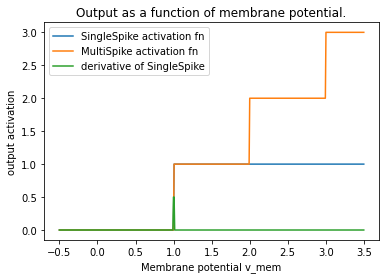
If we replace the derivate of that activation function (green line above) with a surrogate gradient function, we will get better results. Here is a plot of a few of those.
x = torch.linspace(-0.5, 3.5, 500)
plt.plot(x, sina.Heaviside(window=0.5)(v_mem=x, spike_threshold=1.0), label="Heaviside")
plt.plot(x, sina.MultiGaussian()(v_mem=x, spike_threshold=1.0), label="MultiGaussian")
plt.plot(x, sina.SingleExponential()(v_mem=x, spike_threshold=1.0), label="SingleExponential")
plt.xlabel("Membrane potential v_mem")
plt.ylabel("derivative of output activation")
plt.legend();
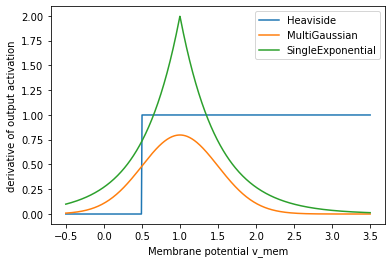
Further up we already defined MultiGaussian as our surrogate gradient function. Let’s see how our new surrogate gradients and surrogate activation function look like.
activations_singlespike.backward(v_mem)
plt.figure()
plt.plot(
v_mem.detach(), activations_singlespike.detach(), label="SingleSpike activation fn"
)
plt.plot(
v_mem.detach(),
np.gradient(activations_singlespike.detach()),
label="derivative of SingleSpike",
)
plt.plot(v_mem.detach(), v_mem.grad.detach(), label="MultiGaussian surrogate deriv.")
plt.title("Single spike activation, derivative and surrogate derivative")
plt.xlabel("Membrane potential v_mem")
plt.legend();
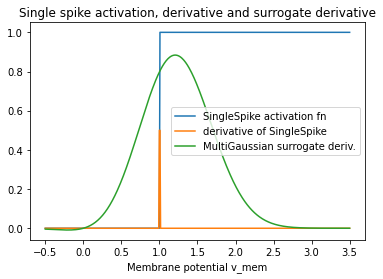
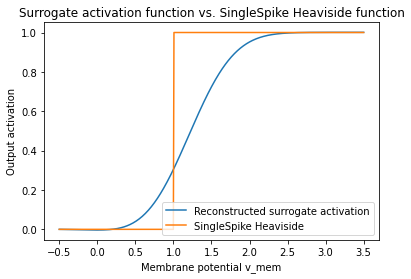
When we accumulate those surrogate gradients, we can reconstruct our surrogate activation function.
surrogate = np.cumsum(
v_mem.grad.detach().flatten() * (v_mem[1] - v_mem[0]).detach().item()
)
plt.plot(v_mem.detach(), surrogate, label="Reconstructed surrogate activation")
plt.plot(
v_mem.detach(), activations_singlespike.detach(), label="SingleSpike Heaviside"
)
plt.title("Surrogate activation function vs. SingleSpike Heaviside function")
plt.xlabel("Membrane potential v_mem")
plt.ylabel("Output activation")
plt.legend();